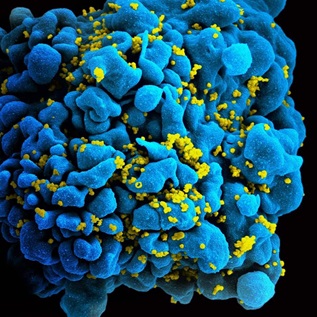
Better Predictions of Flu Strains Could Help Prevent, Lessen Outbreaks
2 Pew scholars explore methods to forecast virus mutations
This year’s flu season has the potential to be the worst in more than a decade. Two Pew supported researchers are working to better predict which strains will be the most dangerous so scientists can develop vaccines accordingly.
iStockphotoBetween 291,000 and 646,000 people worldwide die from seasonal influenza-related respiratory illnesses each year, according to the Centers for Disease Control and Prevention. The flu can be especially dangerous for those 65 and older and children under 5.
The 2017-18 flu season has been among the most brutal in years. Between Oct. 1 and Jan. 20, 11,965 confirmed flu hospitalizations nationwide were reported to the CDC. The wide impact has been attributed in part to the influenza strain now circulating and how well this year’s flu vaccine protects us against it.
In the months leading up to flu season each year, researchers at the World Health Organization identify a few strains they think will infect people in the Northern Hemisphere—based primarily on what made people sick during the Southern Hemisphere’s most recent flu season. Designing an effective vaccine requires an accurate prediction of which strains of the virus will predominate in the coming season—projections that can be challenging to make. Two Pew biomedical scholars at the Fred Hutchinson Cancer Research Center in Seattle are working to combat future flu outbreaks by developing tools to better predict which strains will make people sick each year.
Trevor Bedford, a 2016 Pew scholar, is using computers to model viral mutations in an effort to improve forecasts and predictions to thwart future epidemics, and to inform public health policies. In 2014, Bedford’s lab launched Nextflu, a website designed to track the evolution of the influenza virus in real time. His team has developed novel software to visualize the global migration of the flu virus. Bedford’s approach has the potential to extend to tracking other evolving viruses, such as Ebola, chikungunya, and Zika.
Jesse Bloom, a 2015 Pew scholar, is also trying to understand and predict how viruses evolve. His lab looks into how mutations affect viruses’ ability to replicate and spread. Bloom’s team is generating pools of flu viruses that contain genetically engineered mutations so scientists can determine which of the mutations could support viral replication and which are specifically targeted by the immune system.
Wired magazine wrote about Bloom’s work last year and described a new technique to study how flu viruses evolve—by examining mutant strains in sinus fluids from cancer patients who had long-term infections. Interestingly, Bloom’s team found that mutant strains similar to those taken from these patients ended up spreading globally the following year. The research could offer clues to guide the creation of new flu remedies and provide a basis for predicting which flu strains will evolve and prove most successful at infecting humans, the key to designing an effective seasonal vaccine.
Studies have shown that the annual vaccine typically can reduce the risk of contracting the flu by 40 to 60 percent if circulating strains are predicted accurately and were included in the vaccine. While the effectiveness of the vaccine varies each year, the research that Bedford and Bloom are conducting could drastically improve vaccine effectiveness by helping public health organizations make better-informed decisions on the strains to include in the seasonal variation—and by giving manufactures the time to produce enough doses for the public.
Kara Coleman directs The Pew Charitable Trusts’ biomedical programs, including the biomedical scholars, Pew-Stewart Scholars for Cancer Research, and Latin American fellows programs.