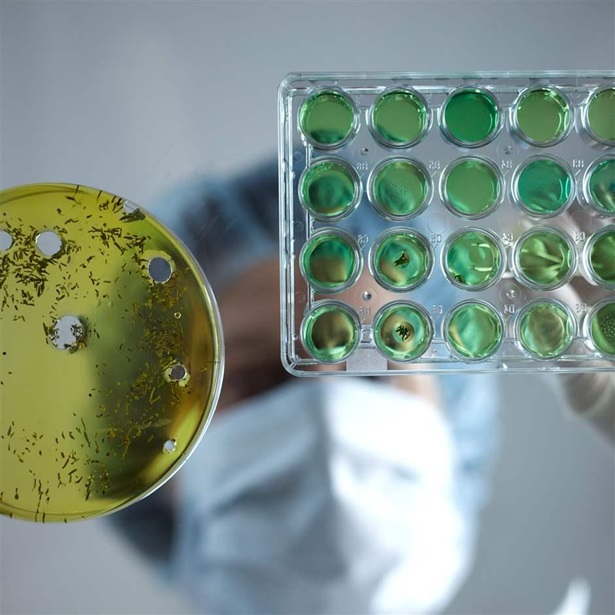
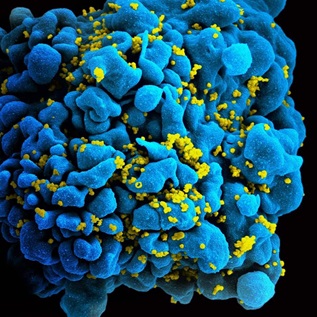
Researcher Looks to Plants in Search for New Antibiotics
Superbug survivor discusses effort to combat antibiotic-resistant bacteria—and why she’s making her data public

Effective November 18, 2021, Pew transferred all SPARK data to The University of Queensland’s Community for Open Antimicrobial Drug Discovery (CO-ADD). Please visit spark.co-add.org or contact [email protected].
Dr. Cassandra Quave’s path to her work as a leader in antibiotic drug discovery research initiatives at Emory University in Atlanta started when she was a child and she and her family dealt with her own serious health issues that have had life-long repercussions.
Now an associate professor of dermatology and human health at the university, Quave also serves as an ambassador for Pew’s Stand Up to Superbugs initiative, which brings together researchers, patients, doctors, farmers, and others to share their stories and expertise about the growing threat of antibiotic-resistant bacteria with policymakers and to urge increased action.
Her latest works—published in Chemical Reviews and Frontiers in Pharmacology—together represent the first comprehensive report to look in depth at the scientific literature on plants with antibacterial properties. She has shared all of this catalogued data via Pew’s Shared Platform for Antibiotic Research and Knowledge (SPARK), where researchers around the world can access the data for free and use it in their own antibiotic discovery efforts. This interview has been edited for clarity and length.
Q: How did you first become aware of the problem of antibiotic-resistant bacteria?
A: I’ve been interested in antibiotic resistance since I was young. I was born with multiple congenital birth defects, which meant I went through a lot of surgeries during my childhood. Doctors had to amputate my leg at the age of 3. Following the surgery, I got a hospital-acquired infection—methicillin-resistant Staphylococcus aureus (MRSA)—that was especially difficult to treat because it had invaded my bone tissue. That resulted in the loss of more of my leg.
A few years later, in elementary school, I fell in love with microbes and I started putting everything I could find under the microscope—and I mean everything—even dog saliva. As I got older, I started competing in science fairs. Most of my projects involved bacteria of some sort. And when I got to college, I studied ethnobotany, which focuses on the relationships between people and plants, including plant-based medicines. Trips to the Amazon during my junior and senior years opened my eyes to how much people around the world still rely on plant-based drugs, and that many drugs in Western medicine—including antibiotics—were originally discovered in nature. By the time I went to graduate school, things came full circle; I combined my love of nature-based medicine with my interest in antibiotic-resistant bacteria.
Q: Can you tell us about your recently published literature review and why you wrote it?
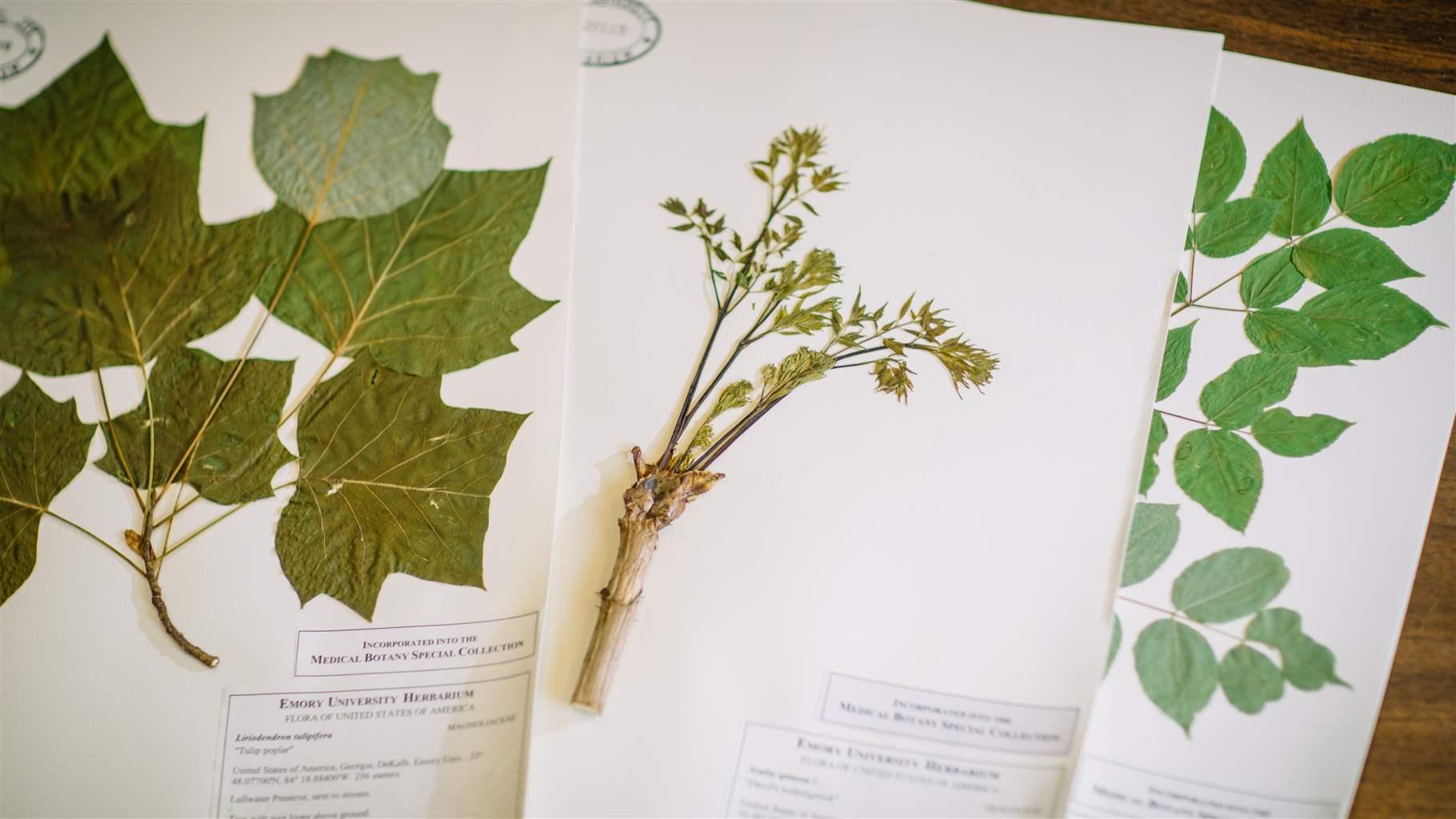
People have used plants to manage infectious diseases throughout human history. Of the 374,000 or so plant species already discovered on Earth, about 28,000 are used in medicine today. However, not many of these plants have been scientifically investigated in depth. We wanted to ask the question: What do we actually know about the antimicrobial potential of these plants in combating bacterial pathogens?
My team and I reviewed the scientific literature related to plants’ antibacterial properties, examining all papers on this topic published from 2012 to 2019. We ultimately focused on 653 of the most relevant and rigorous studies and found data on almost 1,000 plant species and 500 plant-derived compounds that had been evaluated for their antimicrobial potential. It was an exciting opportunity to get a comprehensive look at what the existing research says and where the gaps are. My hope is that our papers on this topic can serve as a guide for more systematic and standardized research of plant products moving forward and help increase awareness among the antibiotic discovery community of the vast potential in the field of plant research.
Q: What do you think is the biggest challenge facing antibiotic discovery right now and how do you believe natural products can address it?
Plants are chemically complex, and while this can present a challenge in the lab, it shouldn’t scare us away from studying plant compounds for antibacterial potential. This complexity contributed to the scientific community’s shift away from natural products in its search for new antibiotics in the 1980s. But today we have tools at our disposal that we didn’t have then—for example, analytical chemistry tools to better understand the activities of and relationships between plant compounds. It seems logical to me that we should investigate sources that have already been used—in some cases for millennia—by humans to treat infections. Instead of being fearful of plants’ complexity, let’s take advantage of it.
Q: What drew you to SPARK as a repository for your review data?
When I first learned about SPARK, I thought it was a great initiative. As researchers in the antibiotic discovery field, we worry about losing the intensive knowledge base and internal datasets from companies when they shut down. Sharing both successes and failures helps scientists avoid repeating failed studies. How can those of us in the field know that a molecule has already been studied if it is not in the published literature and we don’t have access to those datasets? SPARK helps solve this problem.
This is also why we share our plant bioactivity data online via Emory’s herbarium and use other herbarium databases across the globe. When we’re looking for antibacterial properties in new plants, it’s helpful to have access to past data that can accelerate identification of plant molecules that may already be known. This type of data sharing also can help us narrow down the scope of which species to target when searching for new molecules, which allows us to better target our limited research funding and capacity.
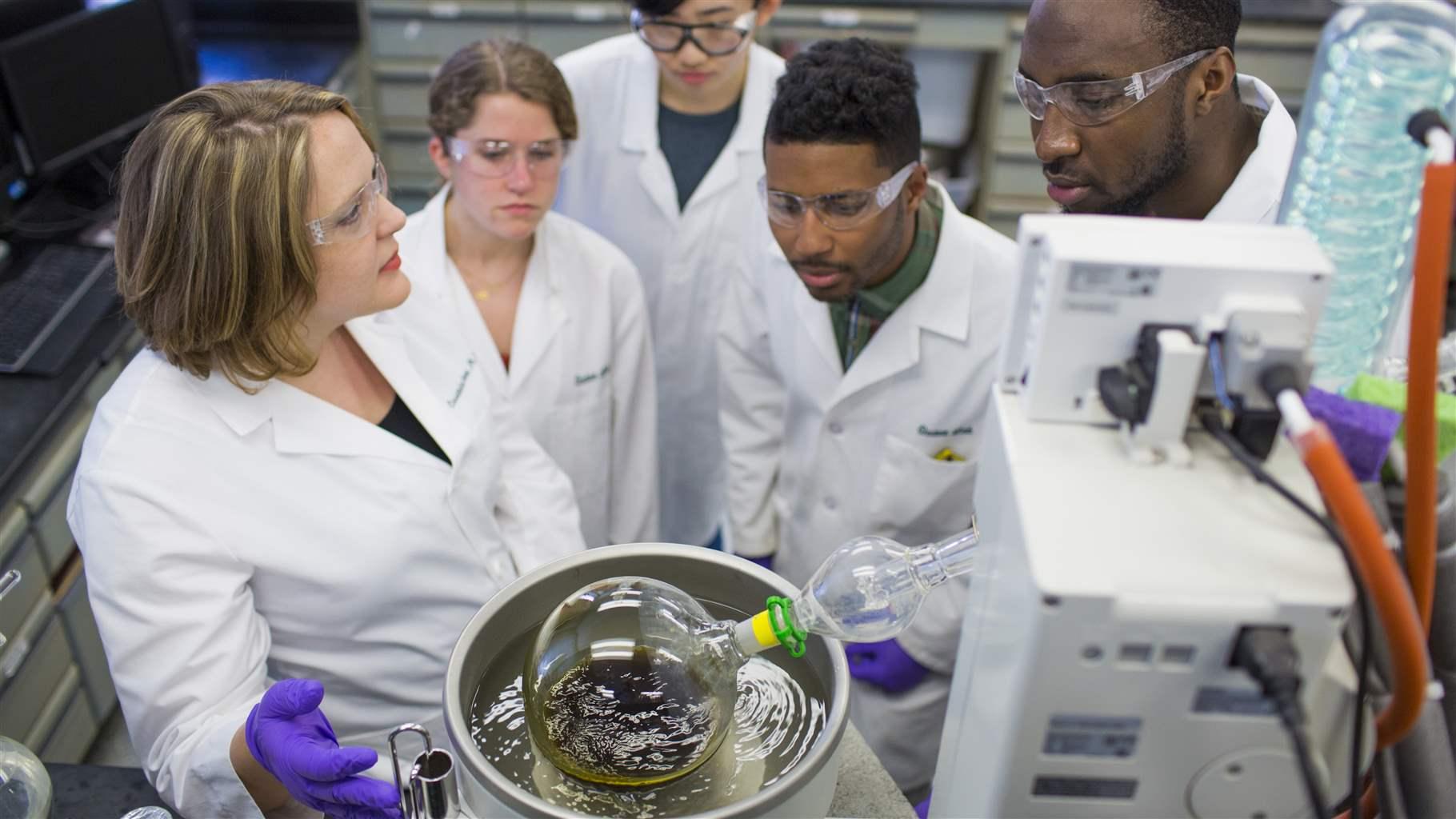
In the case of my new papers, we had a team of 16 researchers—botanists, pharmacologists, microbiologists, and natural product chemists—working for more than a year to curate relevant studies and vet the supporting data. I want the broader antibiotic discovery community to benefit from this work, not spend another year re-creating it.
Q: What do you wish people better understood about the search for new antibiotics?
I wish the general public had a better appreciation for the clear and present danger that superbugs present for health care. I think a lot of people underestimate the impact of antibiotic resistance on the safety of basic procedures such as surgical interventions, childbirth, and cancer therapies.
I’d love to see more of an openness to nontraditional approaches from the scientific community. Our current approach to treating bacterial infections is less than a century old—we can still learn from the other ways that humans have treated these infections for millennia. Keeping an open mind about the impact of natural products on infectious disease might uncover much-needed new solutions.


America’s Overdose Crisis
Sign up for our five-email course explaining the overdose crisis in America, the state of treatment access, and ways to improve care
Sign up
Scientists Need to Develop New Antibiotics Soon