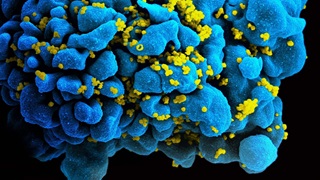
DNA in Seawater Reveals Presence of Sharks—Even When They’re Unseen
Novel method is more efficient than traditional techniques in measuring species diversity
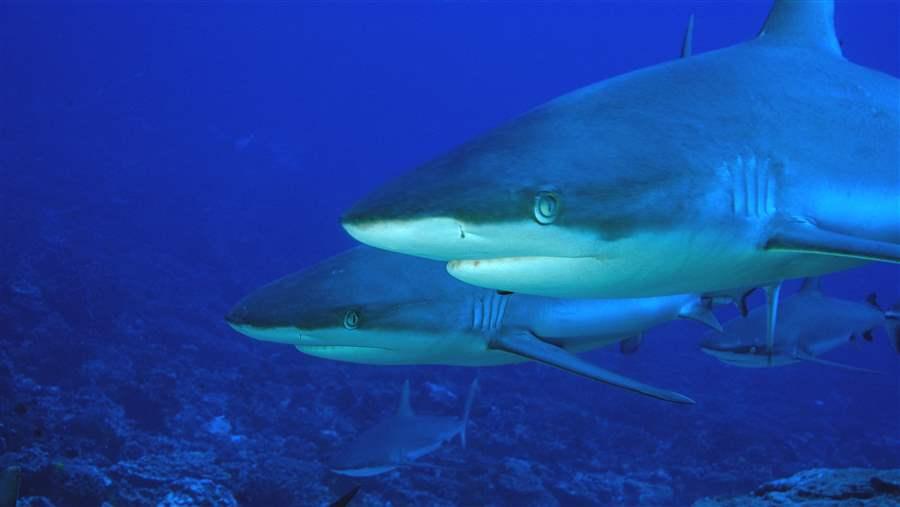

DNA-based research identified 13 species of shark in 22 samples of seawater off New Caledonia, outperforming traditional tracking and identification methods.
Christian GrondinMarine scientists rely on traditional techniques, including underwater video surveys and visual observation, to keep track of migratory species such as sharks. But to do that effectively requires being in the right place at the right time, which is often hard to achieve.
Experts have long sought a way to more easily monitor these highly mobile, elusive, and often rare species. New research shows that the answer just might be environmental DNA, or eDNA.
With sharks facing heavy fishing pressure around the world—at least 63 million and as many as 273 million are killed every year in commercial fisheries globally—policymakers need accurate data to inform effective conservation policies.
With eDNA, scientists use the genetic material left behind by organisms to determine whether they are present in the area. Stefano Mariani, a professor and associate dean of research at the University of Salford, United Kingdom, and a team of researchers based in France, Australia, Polynesia, and the United States conducted traditional and eDNA surveys in New Caledonia, an archipelago in the South Pacific that boasts many shark species. They found that eDNA detected more species than do traditional methods, an indication that the older techniques may be less accurate. The discrepancy was found both in remote areas, where people are less likely to affect sharks, and in regions close to human populations, which the animals may avoid through learned behavior.
Ultimately, having a more accurate picture of shark diversity and richness of species can better inform conservation initiatives and prioritize areas for protection.
“EDNA has great potential for complementing traditional survey methods and improving species detection,” says Mariani. “It could lead to more efficient conservation strategies for threatened species.”
The findings
Of the 26 species of shark known to inhabit the waters of New Caledonia, the eDNA method yielded 13 species from only 22 seawater samples. In comparison, only nine species were observed in 2,758 underwater visual census (UVC) samples, and data from 385 baited remote underwater video stations (BRUVs) also revealed only nine species. Of the 13 species identified using eDNA, six were not identified by either of the two traditional methods, further suggesting that those older techniques are underestimating diversity.
In fact, sharks were identified in 91 percent of the 22 samples taken for eDNA analysis, with two or more species found in 64 percent of these samples. UVCs identified sharks in only 15 percent of samples, and BRUVs identified them in 54 percent. Diversity was also lower using the traditional methods: UVCs detected more than one species in only 3 percent of samples, and BRUVs in only 23 percent.
Advantages and a drawback
EDNA sampling also is much faster than the other techniques. In this study, the eDNA work spanned three months in 2015, while BRUVs took place over three years, from 2012 to 2014, and the UVC surveys took nearly two decades (1986 to 2014). Although eDNA doesn’t reveal the age and size of detected species, it saves scientists the significant time spent on tracking, catching, and tagging and shows promise in helping researchers learn which sharks are where—which would be a major step toward effective conservation policies for these imperiled animals.
Jim Palardy is a project director with The Pew Charitable Trusts’ environmental science division.