Information-Sharing Platform to Fill Knowledge Gaps Impeding Antibiotic Innovation
Answers to common questions about Pew’s publicly available database intended to help scientists build on previous research, generate new insights
This information is out of date. Effective November 18, 2021, Pew transferred all SPARK data to The University of Queensland’s Community for Open Antimicrobial Drug Discovery (CO-ADD). Please visit spark.co-add.org or contact [email protected].
Q. What is SPARK?
A. The Shared Platform for Antibiotic Research and Knowledge is a groundbreaking and dynamic information-sharing platform. It brings together curated antibiotic discovery data and cutting-edge analytics to help scientists tackle the scientific barriers blocking antibiotics discovery. While the global threat of antibiotic resistance continues to rise, nearly every antibiotic in use today is based on a discovery from more than 30 years ago. SPARK will focus on the unique challenges of finding and designing antibiotics that can defeat drug-resistant Gram-negative bacteria, which are among the hardest-to-treat superbugs.
Q. Why did Pew build SPARK?
A. The pipeline of products in development to treat or prevent bacterial infections is stagnant and cannot meet today’s urgent and growing patient needs. Leading experts have repeatedly identified information sharing as essential to spurring innovation; previously, however, there was no publicly available mechanism for sharing data and expertise across the antibiotic discovery research community. As a result, researchers have had difficulty benefiting from the insights of others.
Similar data-sharing tools have helped catalyze drug discovery in other research areas, such as cancer, neglected tropical diseases, and tuberculosis. Pew’s goal is for SPARK to do the same for antibiotic-resistant bacteria, by provide an impetus and a nexus for scientific inquiry in a field desperately in need of a jump-start.
Q. What types of information will be included In SPARK?
A. The information in SPARK is collated and curated by a team of antibiotic discovery experts and includes chemical and biological data relevant to understanding how molecules can get into Gram-negative bacteria and stay there, a critical factor in designing drugs that can defeat these increasingly hard-to-treat pathogens. Currently, the data is pulled from publicly available sources, such as published research articles. However, the platform has the capacity to host and integrate previously unpublished data, as well as prospective research findings from studies still in progress at academic centers and drug companies. For additional detail, see “The Data Driving SPARK.”
Q. Can I share my data on SPARK?
A. SPARK data should be relevant to research into the key scientific barriers to antibiotic discovery and have the potential to enhance scientific understanding of Gram-negative permeability. If you have suggestions on data that should be included in SPARK, or if you are a researcher interested in donating data to the platform, contact us at [email protected]. All data uploads are subject to SPARK’s Terms of Use (donors retain all ownership rights to uploaded data but grant Pew rights to make this data publicly accessible for research and development purposes).
Q. How does SPARK work?
A. Through SPARK’s user-friendly, cloud-based interface, experts from across disciplines (e.g., biologists, chemists, computational scientists) and around the world can work together to uncover new observations, generate and share new hypotheses on how molecules enter and stay inside Gram-negative bacteria, and identify the tools and additional information needed to advance discovery.
SPARK uses technology developed by Collaborative Drug Discovery Inc. (CDD), a software company specializing in cloud-based drug discovery informatics, and is accessed via the CDD website. To optimize the platform, Pew worked with a team of antibiotic discovery experts and engaged researchers in the field to beta test the platform before launch. CDD also provides training and technical assistance to help users and data donors.
Q. Who can use SPARK?
A. SPARK is publicly available for use by researchers around the world across disciplines and sectors—academia, industry, government, and nonprofits—who are focused on antibiotic innovation.
There is no cost to use SPARK. To request a log-in, please email spark-access@pewtrusts.
SPARK creates opportunities to cultivate the next generation of antibiotic discovery researchers and can serve as a valuable resource in overcoming the challenges of “brain drain” and traditionally siloed information sources. This is particularly important given that many antibiotic research programs have been abandoned or downsized in recent years, and findings from previous discovery efforts are often scattered across the academic literature or unpublished.
Q: I received my SPARK log-in information. Now what?
A. To access SPARK, go to the CDD website and enter your log-in information. This will take you directly to the SPARK interface. For an introduction to the platform’s functionality, including information on how to search available data, visit CDD’s “Quick Guide.”
Q: How do I search SPARK?
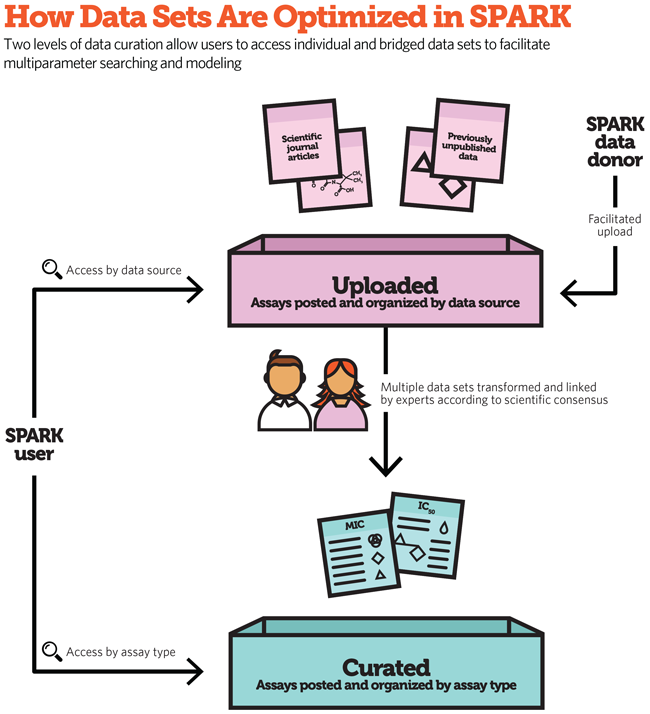
A. Users can search SPARK in various ways, including by compound substructure, keywords, data source, or chemical properties such as molecular weight or pKa, a measure of acidity. The data is organized into two categories, called projects, depending on the type of data, and users can conduct searches within one or both of these data categories:
- Uploaded Data. Contains information that has been organized, validated, and—where applicable—standardized in a way that is easily searchable and usable.
- Curated Data. Includes integrated collections of information pulled from multiple data sets with common elements and comparable variables.
Data collections are being actively populated, so users should check back periodically to see what has been added.
Users can also draw from a Public Data project available on the CDD website, which includes information from other publicly accessible data sets.
For more information on SPARK data and how it can be used, see the following resource:


America’s Overdose Crisis
Sign up for our five-email course explaining the overdose crisis in America, the state of treatment access, and ways to improve care
Sign up
Achaogen Provides Data to SPARK, Pew’s Platform for Antibiotic Discovery Research


The Shared Platform for Antibiotic Research and Knowledge (SPARK)